CALYPSO™
Biomolecular interactions, measured and analyzed with a unique rhythm.CALYPSO
Comprehensive Interaction Analysis.
CALYPSO software is a versatile, unique tool for characterizing a wide range of molecular interactions quickly and reliably using composition-gradient, multi-angle static light scattering (CG-MALS). The CALYPSO software integrates smart method design, simple hardware manipulation, and robust data acquisition for a smooth experimental run. Once the measurement is complete, the intuitive graphical user interface steps the user through data processing for rapid but powerful analysis of macromolecular interactions.
Learn how you can explore the interactions between your macromolecules.
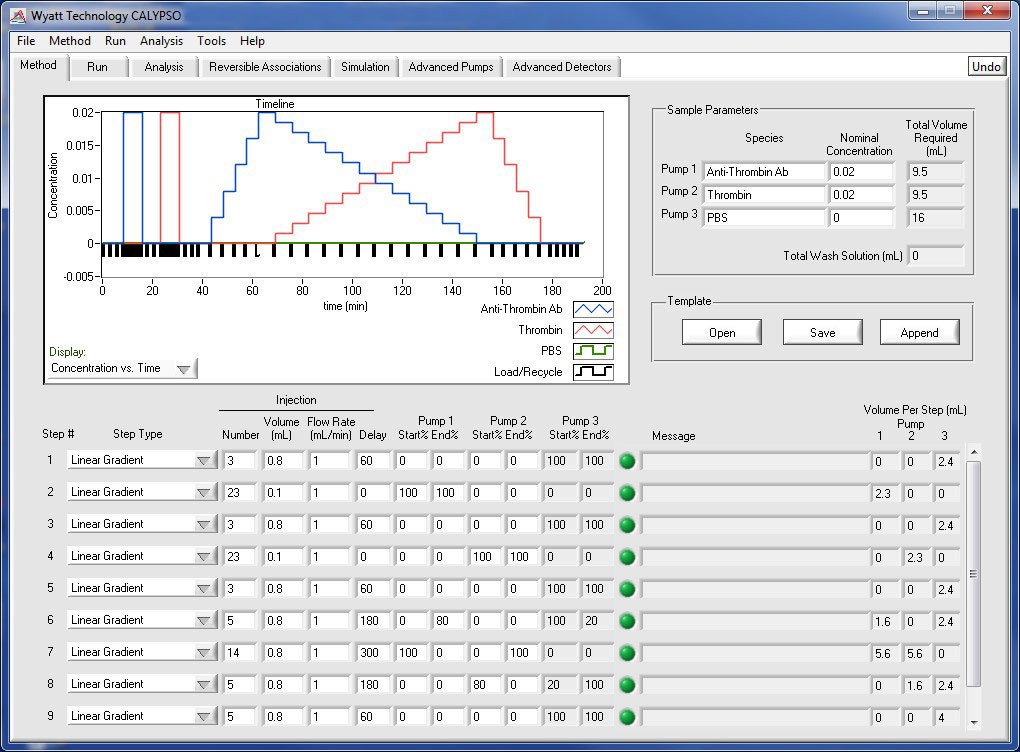
Program comprehensive methods for analyzing self- and hetero-association
Specific, Reversible Interactions – Protein Binding Studies
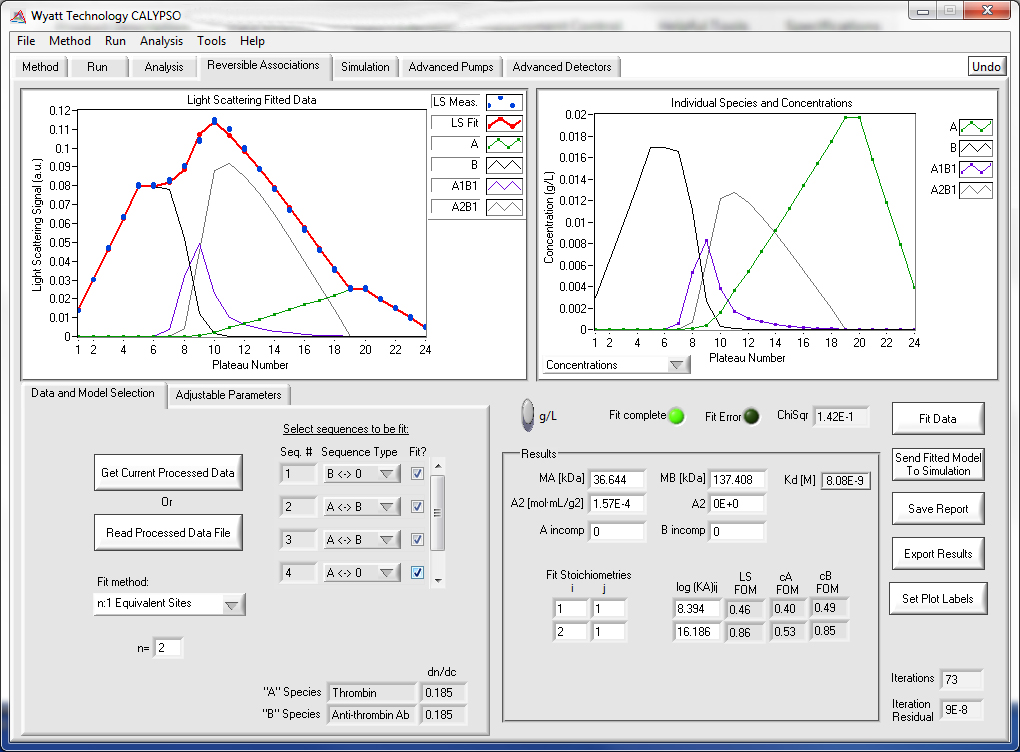
The CALYPSO software uses CG-MALS data to calculate the affinity and stoichiometry of specific, reversible complexes formed in solution, such as antibody-antigen interactions and protein dimerization. The software enables quick data fitting to simple models of dimer formation, 1:n stoichiometries, and infinite self-association. Alternatively, the highly versatile analysis tools allow you to choose and design more complex binding schemes, including simultaneous self- and hetero-association, cooperativity, and meta-complex formation.
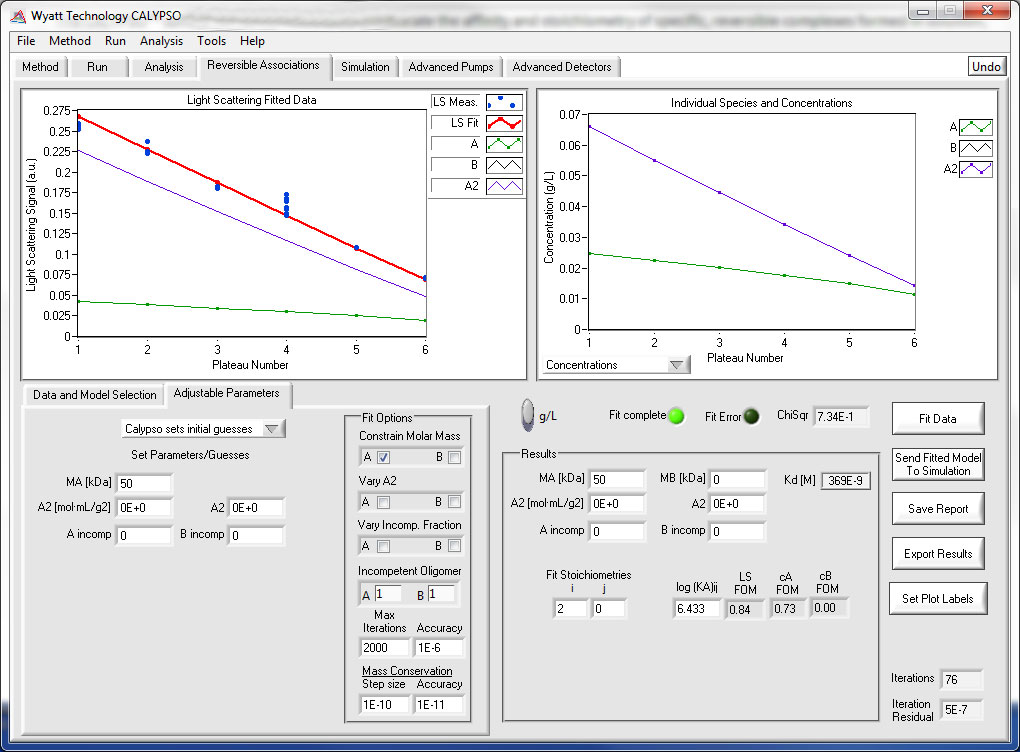
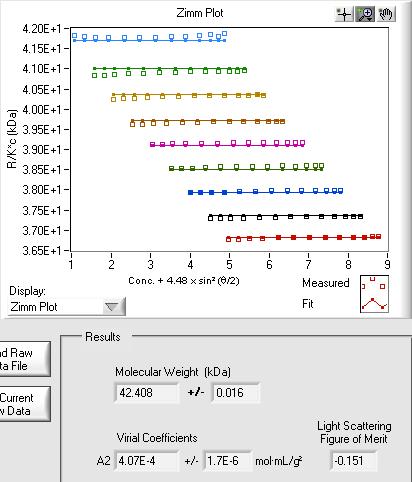
Nonspecific Interactions –Virial Coefficients
The second virial coefficient (A2) is used to quantify nonspecific attractive and repulsive interactions for proteins, polymers, and colloids. Batch light scattering data collected with the CALYPSO software can be analyzed using a Zimm formalism, Debye formalism, or effective hard sphere approximation. Higher virial coefficients may be determined as well.
Time-Dependent MALS – Association and Dissociation Kinetics
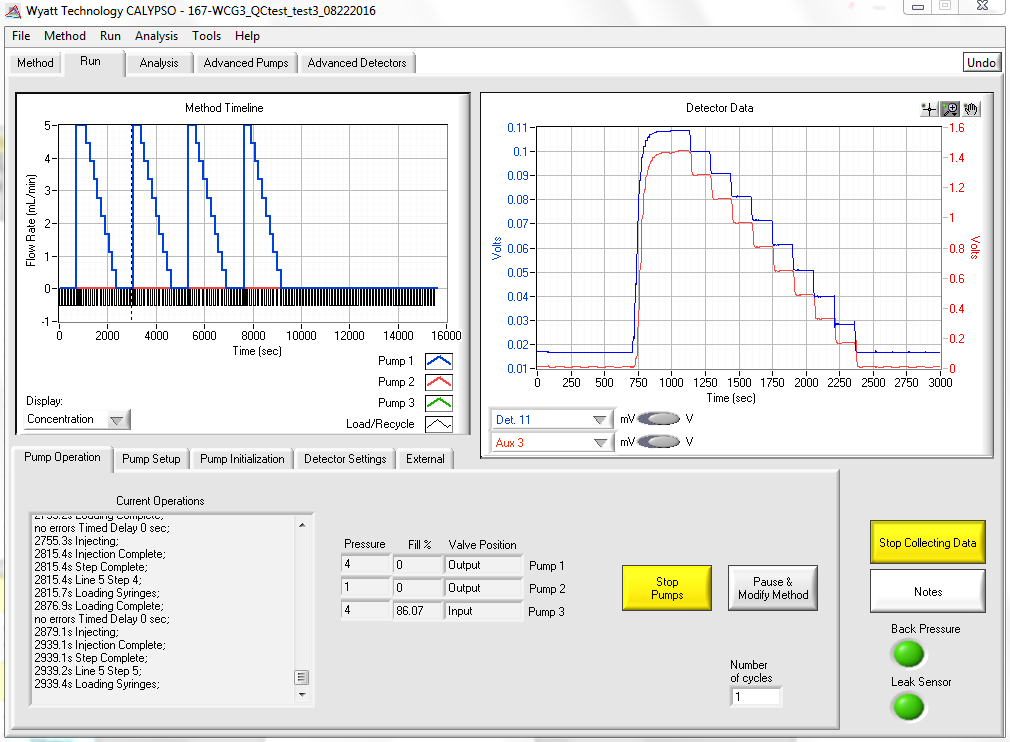
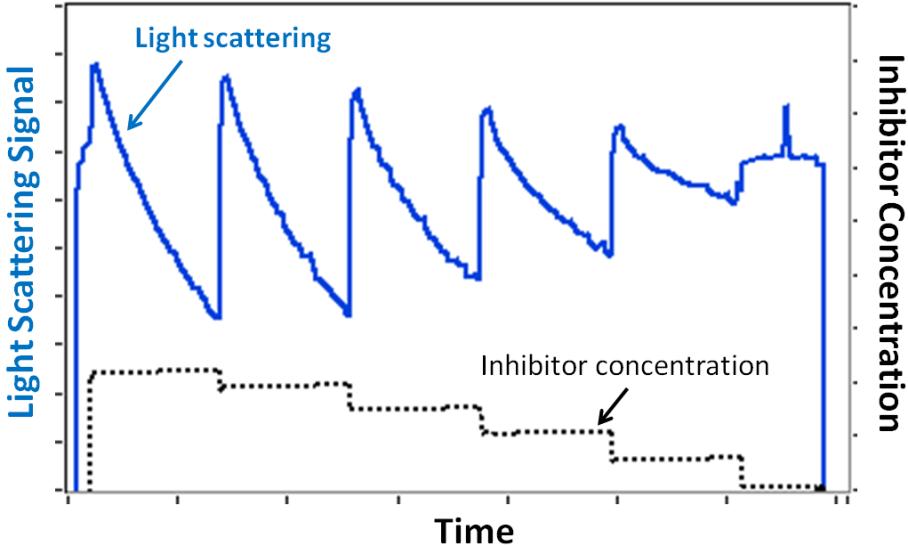
Some protein association or dissociation reactions can take seconds or minutes to reach completion. The CALYPSO software gives you the power to set the time for allowing each composition to come to equilibrium—from a few seconds to several hours. In the example pictured below, different concentrations of a small molecule inhibitor were used to disrupt the dimerization of a protein; the light scattering decrease corresponds to the dissociation of dimers into monomers over time.
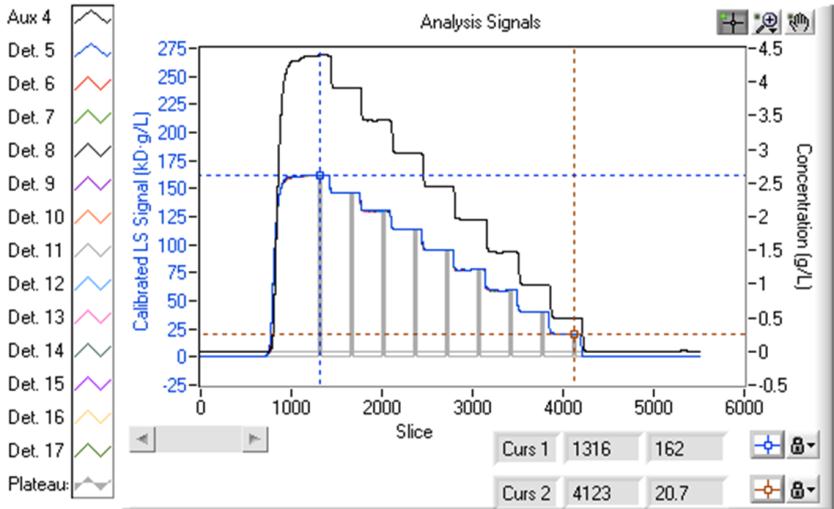
Light Scattering and Concentration Detection
Maximum versatility of experimentation is provided by the real time display of your light scattering and concentration data with a method timeline showing you exactly how far your experiment has run. Analyze data as it is being collected to monitor and evaluate progress of your experiment. Pause and modify your method at any time to optimize and change parameters on the fly, such as increasing measurement times to accommodate long kinetics.
Simultaneous collection with ASTRA
Light scattering using either the DAWN™ or miniDAWN™ and concentration data from a differential refractive index detector (such as the Optilab™) or a UV detector can be collected simultaneously with ASTRA™.
Flexible Concentration Detection
Use either UV or RI concentration detectors or nominal concentrations based on the calculated dilutions. These values are automatically computed and saved with your data. Concentration detection modes can be easily selected post-acquisition.
Calypso Pump Controls
The CALYPSO software allows you to customize every aspect of your CG-MALS experiment, giving you full control of your interacting system. Power on degassers, set maximum allowable flow rates and pressures, and perform priming and wash steps to prepare your hardware for an experiment. Flow and wash parameters can be programmed into an experiment method for complete, unattended experiments.
Useful Accessories
External devices may be readily integrated with the Calypso. The Calypso™ can control external valves to simplify system maintenance. Use an Orbit™ Solvent Recycler to keep clean solvent pumping through your system when not in use. Switch between two wash buffers using an external two-position valve for thorough washing after an experiment. Both functions can be automated within a method for quick and easy cleanup.
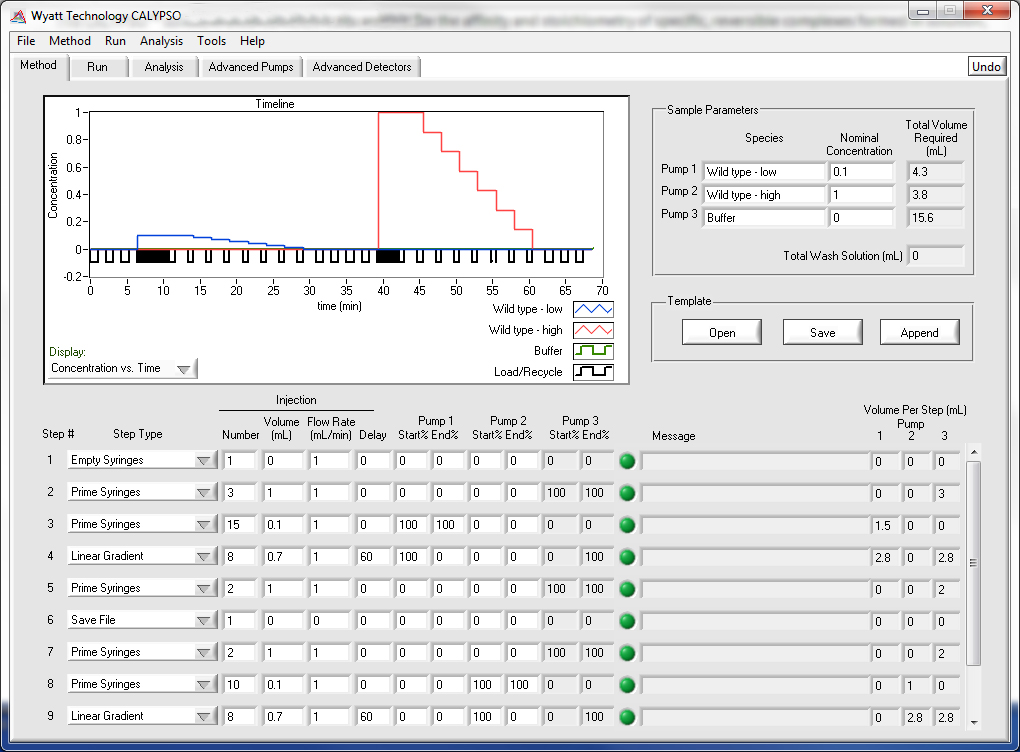
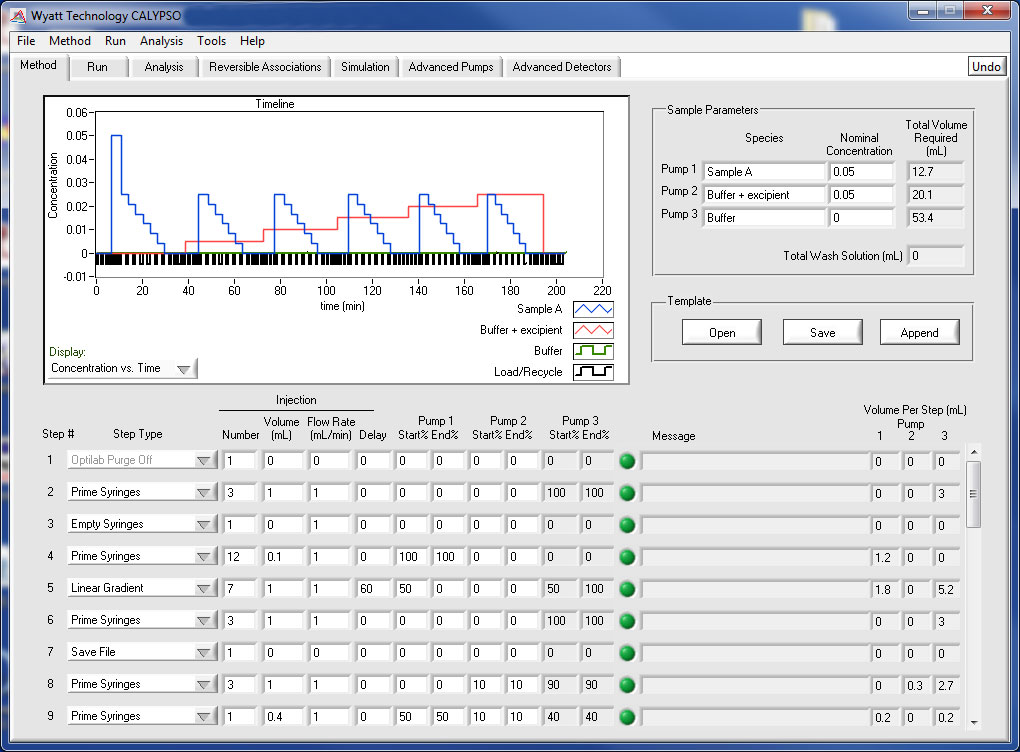
Method Design
Design your own composition gradients or use one of the software's built-in methods to investigate different macromolecular interactions. Any number of composition gradients may be run in a single experiment, repeat runs can be saved to their own data files, and the system washed and ready for the next use in the morning all from a single method page. In addition to the provided example methods, user-designed methods can be saved for later use.
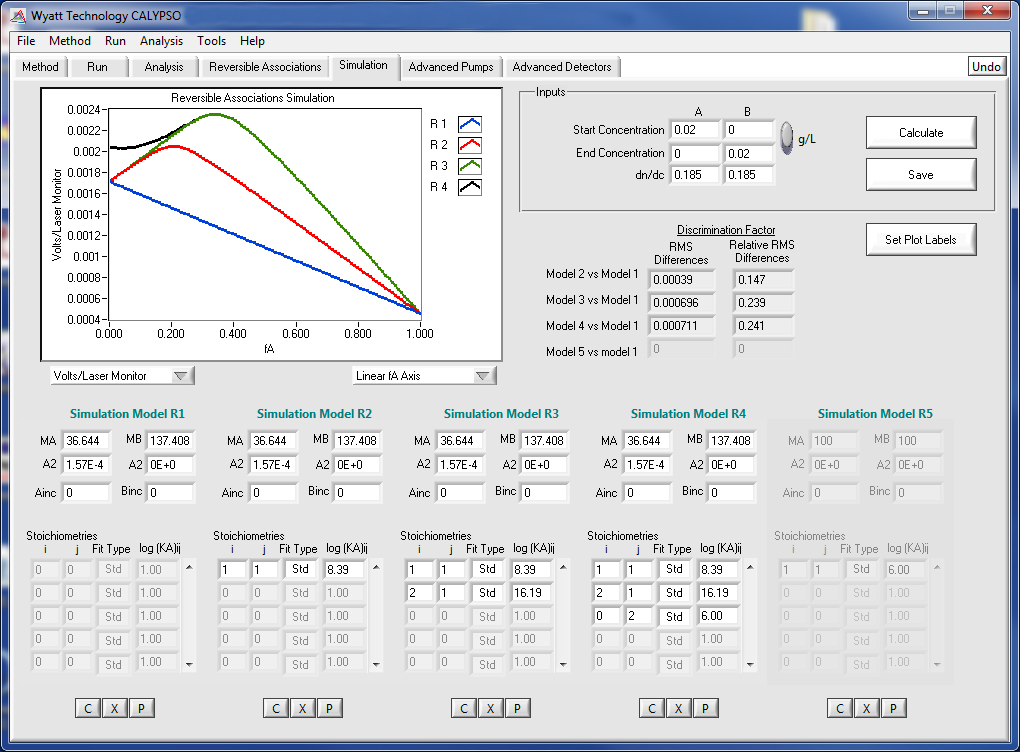
Simulations
Whether you've already performed an experiment and like to optimize parameters or whether you like to hit the ground running designing your first Calypso Experiment, the CALYPSO software provides vital simulation capabilities to aid in experiment design. Input parameters, such as molecular weights, virial coefficients, association stoichiometries, and equilibrium association constants (KA), and the Simulation tool plots predicted light scattering data for up to 5 models. This information lets you determine appropriate solution concentrations for your experiment as well as the optimal composition gradient steps.
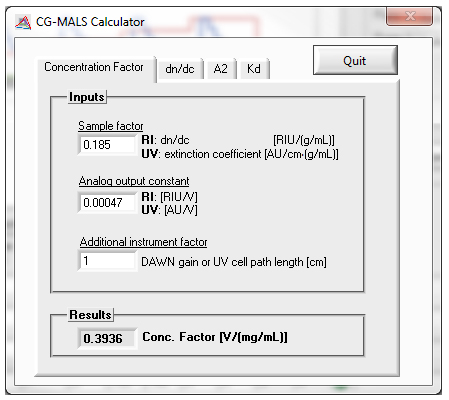
Calculators
Calculators are included in the CALYPSO software to aid in experiment design and evaluation of results: Use the A2 Calculator to predict the excluded volume A2 and select an appropriate concentration range. The dn/dc Calculator helps you estimate the change in your molecule's dn/dc in buffers of varying refractive index. The KD Calculator calculates the equilibrium dissociation constant (KD) for one or two binding sites and helps identify cooperativity.
Host Computer Requirements
Please see Host Computer Requirements for details of the computer hardware and operating system required by CALYPSO.