CG-MALS
Label-free and immobilization-free, CG-MALS is a rigorous yet versatile first-principles technique that measures interactions directly by monitoring the changes in solution molar mass that arise from the formation or dissociation of complexes. Apply CG-MALS to characterize:
- Self- and hetero-association
- Binding affinity from pM to mM
- Absolute molecular stoichiometry (not just mole ratios)
- Multi-valent or cooperative interactions; simultaneous self- and hetero-association
- Kinetics of binding, aggregation or dissociation over time scales of seconds to hours
CG-MALS characterizes self- and hetero-association.
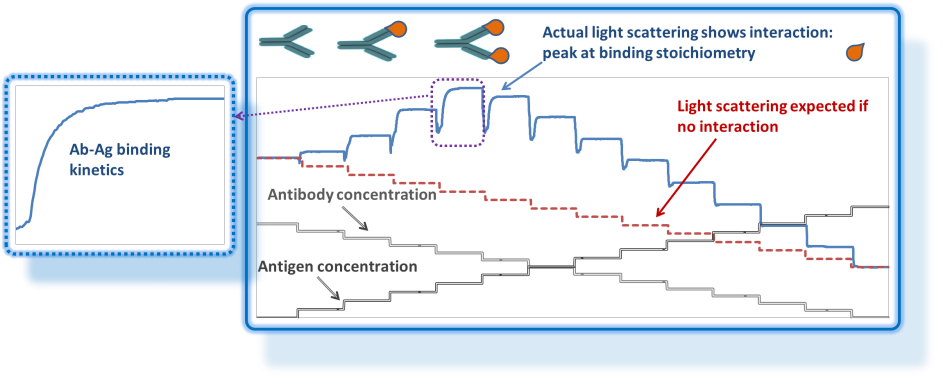
CG-MALS works by mixing different compositions of samples and diluents, then measuring the weight-average molar mass of the solution at each composition step. Binding kinetics are often apparent.
CG-MALS also characterizes non-specific interactions between proteins at high concentrations and between proteins and excipients, which impact colloidal stability of formulated biotherapeutics. Contact Wyatt to learn how CG-MALS can address your specific biomolecular interaction analyses, such as:
- Drug-target binding for monoclonal antibodies or bispecific fusion proteins
- Monovalent or multi-valent receptors
- Self-association under a variety of buffer conditions
- Multi-protein assemblies in structural biology
- Colloidal stability at protein concentrations of 100 mg/mL and beyond
CG-MALS excels at analyzing multi-protein assemblies.
Application notes
Selected references
Attri, A. K.; Fernández, C.; Minton, A. P. Self-association of Zn-insulin at neutral pH: investigation by concentration gradient static and dynamic light scattering. Biophys. Chem. 2010, 148, 23-27.
Esfandiary, R.; Hayes, D. B.; Parupudi, A.; Casas-Finet, J.; Bai, S.; Samra, H. S.; Shah, A. U.; Sathish, H. A. A systematic multitechnique approach for detection and characterization of reversible self-association during formulation development of therapeutic antibodies. J. Pharm. Sci. 2013, 102, 3089-3099.
Monterroso, B.; Alfonso, C.; Zorilla, S.; Rivas, G. Combined analytical ultracentrifugation, light scattering and fluorescence spectroscopy studies on the functional associations of the bacterial division FtsZ protein. Methods 2013, 59, 349-362.
Smith, M. H.; Lyon, L. A. Tunable encapsulation of proteins within charged microgels. Macromolecules 2011, 44, 8154-8160.
Some, D. Light scattering based analysis of biomolecular interactions. Biophys. Rev. 2013, 5, 147-158.
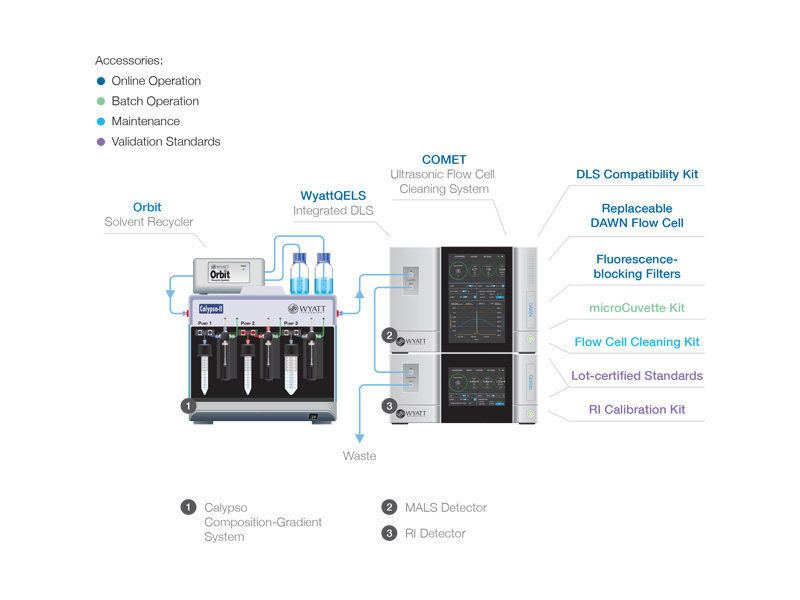
Instrumentation for CG-MALS
Composition-Gradient System
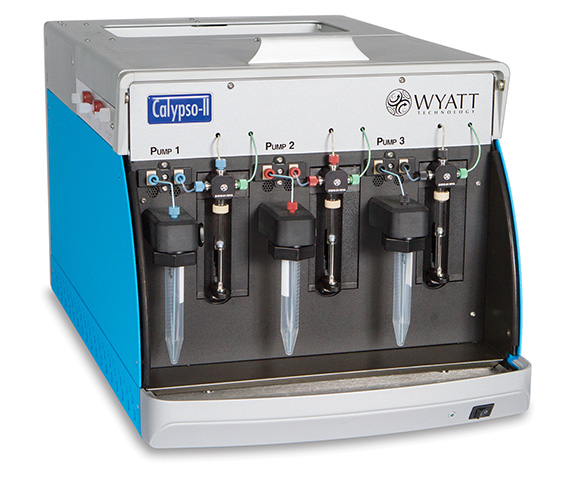

Calypso™ - Composition-gradient preparative system + software for carrying out and analyzing CG-MALS interaction analyses. Requires a MALS detector.
MALS Detectors

DAWN™ - The most sensitive MALS detector available, anywhere. Incorporates detectors at 18 angles to determine molar masses from 200 Da to 1 GDa and radii from 10 – 500 nm.
- Standard option: ambient temperature
- Heated/cooled option: -15 °C to +150 °C
- High-temperature option: ambient to +210 °C
The DAWN offers special options to handle fluorescent samples: fluorescence-blocking filters and an infrared, 785 nm laser.
miniDAWN™ - Second only to the DAWN in sensitivity. Incorporates detectors at 3 angles to determine molar
masses from 200 Da to 10 MDa and radii from 10 – 50 nm. Ambient only.
DynaPro™ NanoStar™ - With sample volumes as small as 1.25 µL and temperature control spanning -15 °C to +150 °C, the NanoStar goes above and beyond traditional cuvette-based DLS instruments: it offers an optimized static light scattering detector (in parallel to the DLS detection system) which may be used with the CALYPSO™ software to perform CG-MALS experiments with very little sample. Samples must be prepared manually, without benefit of the Calypso's automation.
Dynamic Light Scattering Detector
The CALYPSO™ software does not analyze DLS data, but can be set to trigger DLS data acquisition by Wyatt’s ASTRA™ software.
WyattQELS™ - A dynamic light scattering (DLS) module which integrates into a DAWN or miniDAWN MALS detector to provide simultaneous DLS measurements in the same scattering volume.
DynaPro™ NanoStar™ - With sample volumes as small as 2 µL and temperature control spanning -10 °C to +120 °C, the NanoStar goes above and beyond traditional cuvette-based DLS instruments: it offers an optimized static light scattering detector (in parallel to the DLS detection system) which may be used with the CALYPSO software to perform CG-MALS experiments with very little sample. The NanoStar does double duty as an online DLS detector by installing its optical collection fiber into a Wyatt MALS detector.
Refractive Index Detector

Optilab™ - A unique on-line differential refractometer for measuring concentration of any macromolecule, regardless of chromophores. The high-concentration Optilab accommodates protein concentration up to 180 mg/mL.
Software
CALYPSO™ - The CALYPSO software orchestrates CG-MALS analysis of biomolecular interactions, controlling sample preparation and delivery, data acquisition and data analysis. Provides the most extensive suite of association models available to determine the affinity and absolute molecular stoichiometry of macromolecular interactions including protein-protein binding, binding of aptamers and peptides or proteins, etc., as well as first-order analysis of reaction kinetics.
Interested in other techniques or combining other techniques with CG-MALS?
SEC-MALS: Standard in protein, biopolymer and synthetic polymer characterizations labs around the world, Wyatt MALS detectors are valued for reliable and robust measurements.
FFF-MALS: Coupling an FFF system to a set of Wyatt MALS and/or DLS detectors creates a powerful system for accurate and robust characterization of molar mass and size distributions for simple or complex samples.
DLS: Measure the translational diffusion coefficients Dt of nanoparticles and colloids in solution by quantifying dynamic fluctuations in scattered light. DLS is suitable for ensemble measurements ranging from Rh values of 0.2 nm up to 5,000 nm.
MP-PALS: Wyatt Technology's breakthrough Massively Parallel Phase Analysis Light Scattering (MP-PALS) extends robust ELS measurements to proteins and other biomolecules in native buffer solutions. MP-PALS does everything conventional zeta potential instruments can do and much more.
SEC-IV: SEC-IV may be the optimal means of characterizing materials not amenable to SEC-MALS analysis. By simply adding a DAWN, transform a basic SEC-IV setup into a powerful SEC-MALS-IV polymer characterization station, to analyze absolute determination of molar mass and size regardless of conformation, branched polymers, copolymers and measure Mark-Houwink coefficients ab initio.