Calypso™
Product Overview
Product Description
Brochure
CG-MALS
Technology
Configurations
Detectors
Software
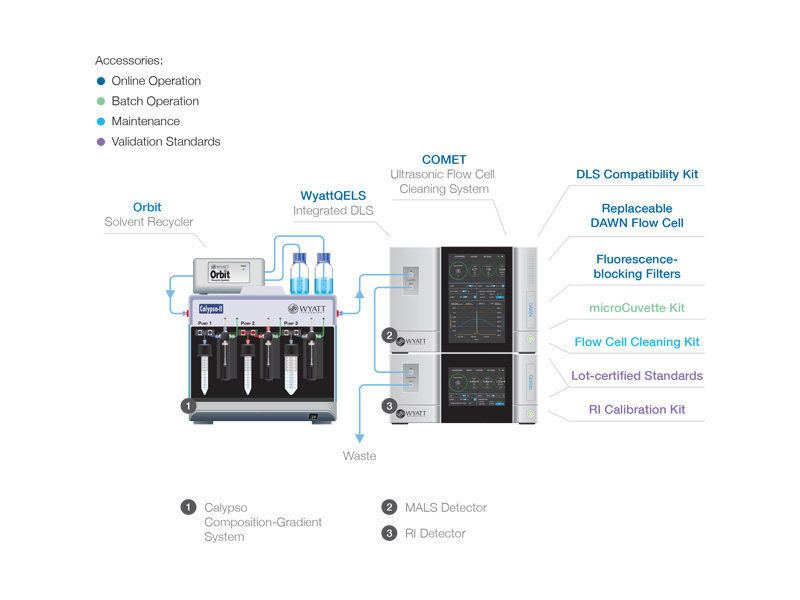
Accessories
Specifications
Publications
Request Info
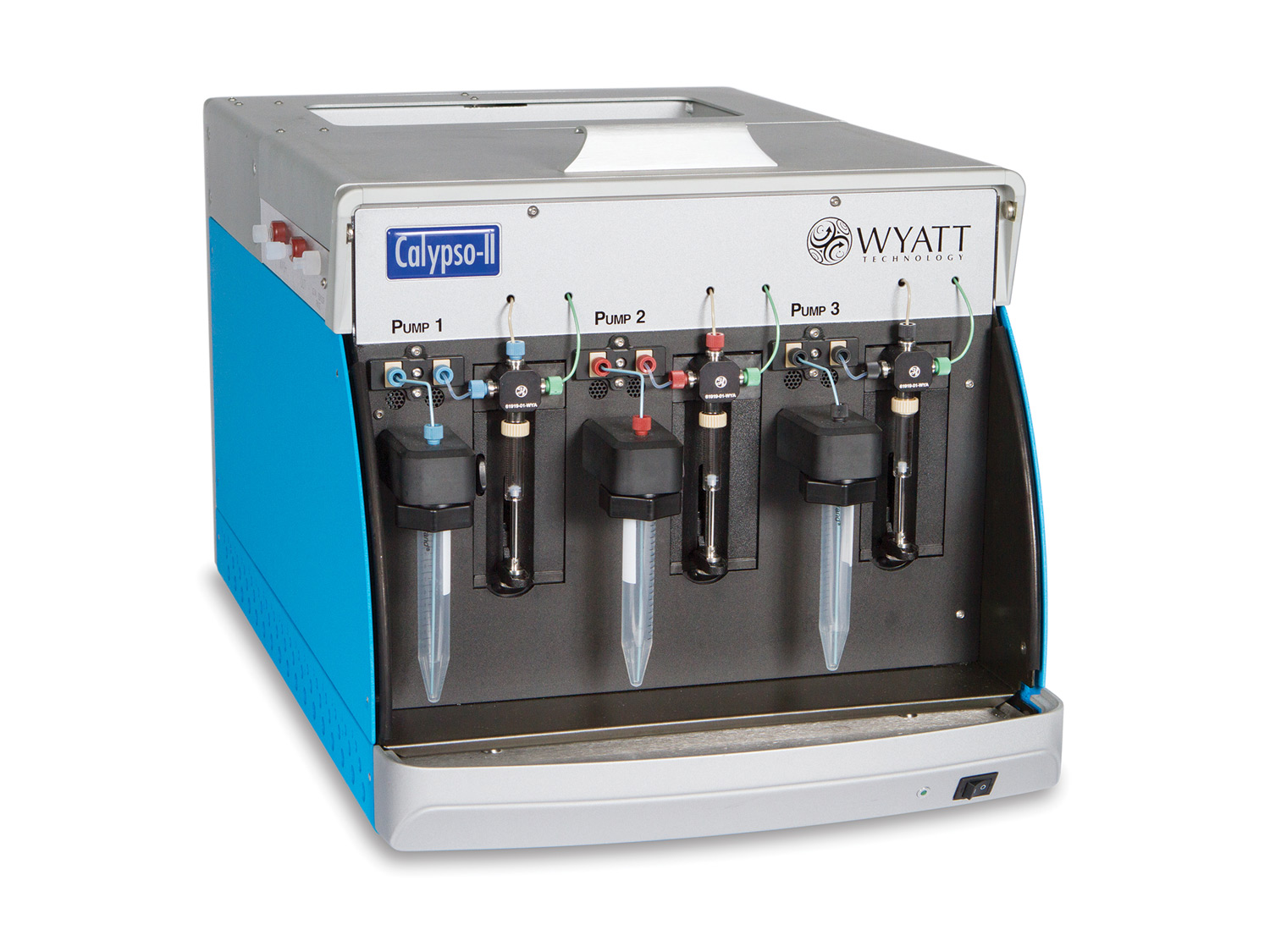

Label-free, immobilization-free characterization of protein-protein and other macromolecular interactions with composition-gradient multi-angle light scattering.
CG-MALS
The Calypso II, in conjunction with a DAWN™ or miniDAWN™ MALS detector, measures binding affinities and absolute, molecular stoichiometries of complex biomolecular interactions.
Weight-Average Molar Mass
In addition to equilibrium association constants Kd and absolute stoichiometry of reversible self- and hetero-associations, it can also help determine reaction or aggregation rates, non-specific interaction parameters (virial coefficients) and even automate measurements of weight-average molar mass and dn/dc for polymers.
CALYPSO™ Software
Included with each Calypso is a copy of CALYPSO software, the most versatile software package available for analysis of biomolecular interactions in solution by light scattering.
Creating composition gradients...
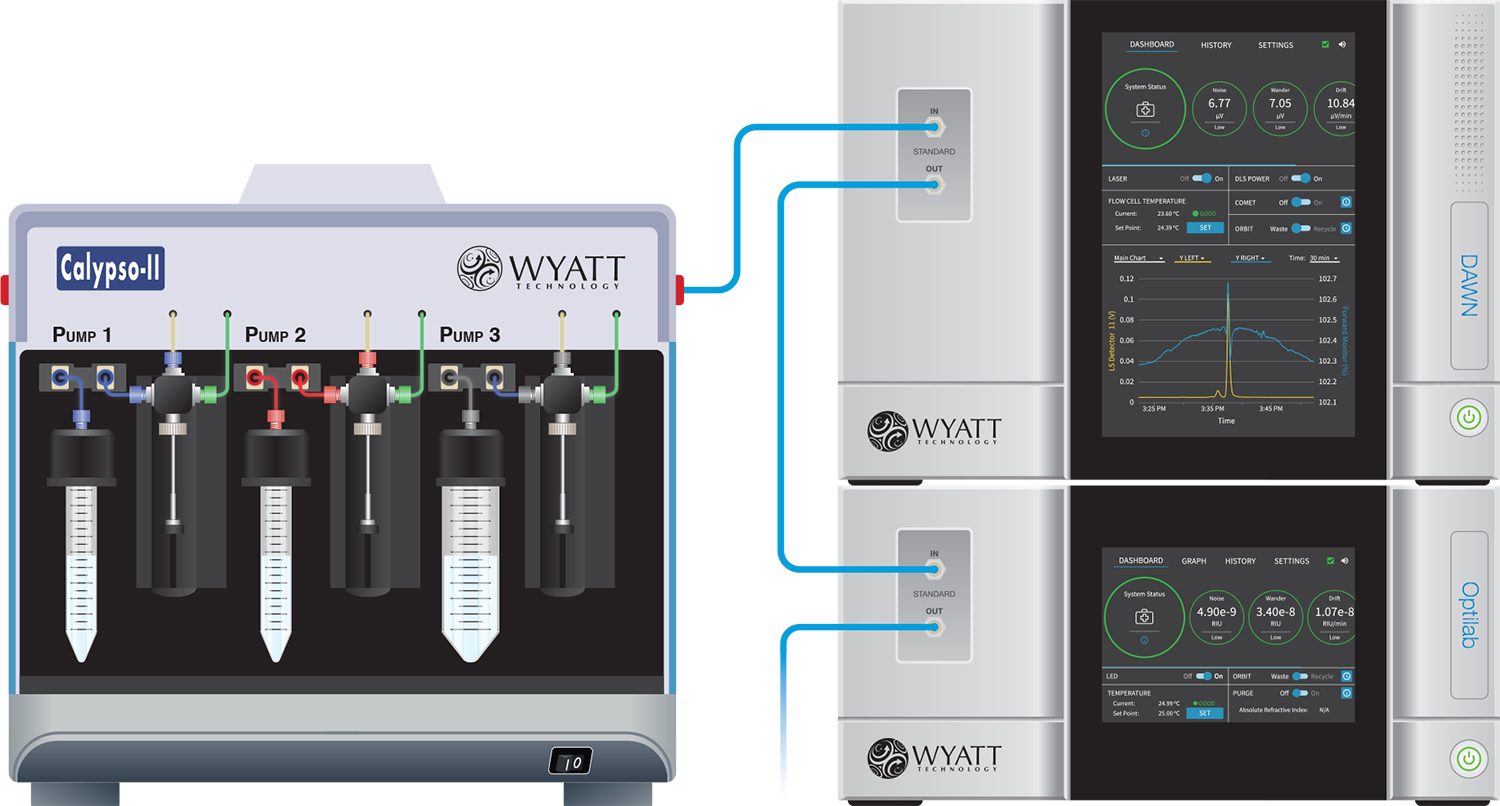
CG-MALS employs a Calypso connected to a MALS instrument and optional concentration detector. The Calypso performs sample preparation and delivery, combining up to three different solution in precise mixing ratios and injecting into the detectors.
Accurate compositions result from setting the Calypso's pumps to appropriate flow rate ratios, all controlled from the CALYPSO™ software through an intuitive GUI. Mixing is achieved by dispensing the solutions simultaneously from all three pumps via a static mixer.
...that work with MALS detectors
Reliable light scattering measurement require the samples to be degassed and filtered. The Calypso incorporates built-in, low-volume degassers and filter housings, connected via PEEK tubing and finger-tight fittings. Even the filter housing is finger-tight, making use of a proprietary design for maximum convenience.
Calypso features
- Biocompatible wetted materials.
- Flexibility to re-organize fluid paths and utilize different wetted materials as needed.
- Convenient loading of sample and diluent solutions via conical tubes: 5, 15 or 50 mL.
- Autoinject Out contact closure signal, to trigger data acquisition by ASTRA™ or DYNAMICS™ software for data acquisition and analyses not supported by the CALYPSO software (e.g., DLS acquisition).
- Built-in Wash port, allowing the pumps to draw on reservoirs of wash solutions for post-experimental clean-up. Automation of up to two sequential wash solutions, e.g. buffer (to clean out proteins) followed by water (to clean out buffer salts) or detergent followed by water, is possible when an Orbit™ solvent selector valve is added to the setup. These operations are appended to the Method for unattended operation.

Absolute characterization of macromolecular interactions
Click here to request a copy of our Calypso brochure.
Label-free, immobilization-free
Composition-Gradient Multi-Angle Light Scattering (CG-MALS) employs a series of unfractionated samples of different composition or concentration in order to characterize a wide range of macromolecular interactions. No special modifications (e.g., sample tagging or immobilization procedures) are necessary: samples are unlabeled and entirely in solution.
Addressing a host of biomolecular phenomena
The primary analysis techniques supported by Calypso hardware and software are:
- Dynamic equilibria: specific binding and complex assemblies of proteins, oligonucleotides and other biomolecules; KD (equilibrium dissociation constant) from picomolars to millimolars; absolute molecular stoichiometry of associating complexes; self and/or heteroassociations
- Non-specific macromolecular interactions: self- and cross-virial coefficients
- Stop flow kinetics: aggregation, dissociation and other time-dependent reactions: equilibration time from seconds to hours
- Zimm plots: concentration gradients for determining solution-average molar mass MW, size Rg, and second virial coefficients
- Refractive index increment: dn/dc
Calypso's automation enhances productivity while the CALYPSO™ software provides an unparalleled selection of interaction models. Relative to other techniques for characterizing protein interactions, such as surface plasmon resonance, sedimentation equilibrium, kinetic exclusion or isothermal titration calorimetry – as well as manual CG-MALS measurements – the Calypso provides fast and accurate results.
Applications
Drug Discovery
- Quantify binding affinity and stoichiometry of enzyme/inhibitor or antibody/antigen interactions, including complex multi-valent and multi-protein complex formation
- Study the impact of small molecules on protein-protein interactions
Process Improvement
- Determine second virial coefficient and adjust buffer parameters to improve formulation stability and viscosity
- Determine cross virial coefficients to optimize antibody purification and understand the effects of large excipients on formulations
Self-assembly / Aggregation
- Quantify impact of solvent ionic strength, pH, or excipients on polymerization or protein associations
- Measure kinetics of self-assembly and aggregation via rate of change of molar mass and radius of gyration, and hydrodynamic radius (with a WyattQELS™ module or NanoStar™ and parallel analysis in ASTRA™ or DYNAMICS™)
Biotech R&D
- Characterize macromolecular binding affinity and associated complex stoichiometry over a wide range of buffer compositions, time, and temperature scales
Deceptively simple
Conceptually, the Calypso is 'just' a box with some syringe pumps and tubing, right? Well, that's what we thought it would be - until coming up against the juxtaposition of challenging biomolecular interaction research and the demands of exquisitely sensitive light scattering instruments. In order to simultaneously achieve:
- High precision
- Minimal sample consumption
- Compatibility from peptides to to high-concentration monoclonal antibodies to nanogels
- Affinities ranging from pM to mM
while still providing exceptional ease-of-use, accuracy, robustness and reasonable cost in capital and consumables, we had to stretch the limits of available technology.
We worked hard, so you don't have to
We left no stone unturned looking for the rare components that met all our demands, such as one-of-a-kind syringe pumps and pressure transducers as well as hard-to-find, 30 nm pore size, filter membranes and low-volume, high-efficiency degassers. What we couldn't buy, we innovated and designed from scratch.
Users have been particularly pleased with our unique, ultra-low-dead-volume, finger-tight filter housings, which help ensure sufficient solution cleanliness for high-affinity analyses yet retain very little sample. Special valves and syringes were co-developed with suppliers to push syringe pump performance and ease-of-use to the max.
Recognizing that CG-MALS is unfamiliar territory for most users, we've also developed Tutorials, Application Guides, Method Templates, and additional support literature and training to help you get started and keep you on track. Make the most of this amazingly powerful technique - we're here to help at every turn.
Some additional Calypso features
The layout and design were fine-tuned to reduce sample use while enhancing automation and maintaining compatibility with the DAWN™ and miniDAWN™, the ultimate in MALS detectors. Special features of the Calypso include:
- Most tubing is 0.010" (0.25 mm) i.d. PEEK for biocompatibility and low volume. May be changed to PTFE or stainless steel for non-aqueous solutions, and larger i.d. for viscous solutions
- A range of syringe sizes to meet specific needs
- In-line degassing of samples and buffers
- Easy access to all tubing and filters, for rapid maintenance
- Pressure warnings indicate when a filter membrane is saturated or a capillary tube clogged
Protein-Ligand Binding
Most protein-ligand and similar biomolecular interactions are best measured at or below 1 mg/mL. The CG-MALS configuration should include:
- Calypso
- DAWN™ H/C for temperature regulation of the interaction and highest sensitivity, though a miniDAWN™ (ambient only) may suffice
- Third-party on-line UV/Vis absorption detector - please Contact Support for recommended models
Non-Specific Interactions
Non-specific interactions are usually measured in the range of 1 - 25 mg/mL. The configuration should include:
- Calypso
- DAWN H/C when measuring temperature-sensitive samples or polymers with Rg > 40 nm; otherwise miniDAWN
- Optilab™ for concentration measurements
High-Concentration Protein-Protein Interactions
These measurements may go to concentrations of hundreds of mg/mL protein.
- Calypso
- DAWN H/C when measuring temperature-sensitive samples or polymers/aggregates with Rg > 40 nm; otherwise miniDAWN
- Optilab HC (high-concentration option) for concentration measurements
- microCuvette™ for concentrated, viscous solutions that cannot be run through the Calypso and MALS flow cell
MALS
DAWN™ - The most sensitive MALS detector available, anywhere. Incorporates detectors at 18 angles to determine molar masses from 200 Da to 1 GDa and radii from 10 to 500 nm.
- Heated/cooled (H/C) option -15 °C to +150 °C
The DAWN offers advanced options such as fluorescence or polarization filters and a 785 nm laser.
miniDAWN™ - Second only to the DAWN in sensitivity. Incorporates detectors at 3 angles to determine molar masses from 200 Da to 20 MDa and radii from 10 to 50 nm. Ambient only.
Options
WyattQELS™ - A dynamic light scattering (DLS) module which integrates inside the DAWN or miniDAWN to provide simultaneous DLS measurements in the same scattering volume. The optical fiber replaces any one MALS detection angle.
Accessories
Orbit™ - The Orbit helps conserve mobile phase by programmatically directing the solvent to either a waste bottle or back to the solvent reservoir. In conjunction with the Calypso, the Orbit is used to programmatically choose from one of two Wash solutions in order to enhance automated post-experimental clean-up.
dRI
Optilab™ - A unique on-line differential refractometer for measuring concentration of any macromolecule, regardless of chromophores. The high-concentration Optilab accommodates protein concentration up to 180 mg/mL.
CALYPSO™ - Simulation and method design of CG-MALS experiments; Calypso fluidics control & MALS/dRI data acquisition; data analysis for determination of equilibrium dissociation constants Kd, absolute molecular stoichiometry and other parameters.
| Syringe Pumps |
3 computer-controlled pumps; dc-servo motor, for minimum pulsation; 3-port distribution valve on each pump for Input, Output and Wash |
| Syringe Volumes | 12.5 µL to 2.5 mL (1 mL and 0.5 mL syringes supplied as standard) |
| Degasser Channels | 1 per pump, 60 µL internal volume |
| Pressure Limit |
1 mL syringes: 200 psi 0.5 mL syringes: 400 psi |
| Auxiliary Inputs/Outputs | Autoinject out, injector valve control, recycle valve control, digital and analog I/O for specific applications |
| Solvents | Aqueous |
| Wetted Materials | Borosilicate glass, PEEK, PTFE, polyethylene, alumina, titanium, stainless steel |
| Typical Sample Volume Per Analysis | 2.5 to 7 mL, depending on detector configuration and complexity of analysis |
| Range of Measurement |
Equilibrium Dissociation Constant, KD: 100 pM to 1 mM typical for 100 kDa molecules (actual range varies with molecular weight and association stoichiometry) |
| Repeatability of Measurement |
Mw: ± 5% |
| Analyses | |
| Specific Binding |
See CALYPSO™ Software for more details.
|
| Non-Specific Interactions |
|
| Kinetics |
1st-order analyzed in software; export data for higher-order kinetics |
| Additional analyses | dn/dc |
| Host PC Requirements |
USB port for Calypso Ethernet connection for DAWN™ / miniDAWN™ communications |
| Dimensions | 58 cm (L) x 37 cm (W) x 35 cm (H) |
Host PC requirements may be found in Computer Requirements.
Specifications subject to change without notice.